Diana Ruggiero is PhD candidate in the Department of Botany and Plant Pathology at Oregon State University. She is researching the genetics of plant development and is using both molecular and computational approaches.
Ms. Ruggiero is currently a USDA-NIFA Predoctoral Fellow through the AFRI Education and Workforce Development program. She previously represented the ARCS Foundation as a supported graduate scholar of the Oregon Chapter.
Before starting graduate school, Diana worked for several years doing ecological restoration, landscaping, and trail work in the Pacific Northwest. She is an AmeriCorps alumna and has a B.A. in Computer Science from Bard College.
You can contact her at either:
ruggiedi@oregonstate.edu
druggiero@protonmail.com
This website was last updated March 14th, 2026.





Education
Oregon State University, College of Agricultural Sciences | Corvallis, ORBard College | Annandale-on-Hudson, NY
Concentration: Experimental Humanities
Publications
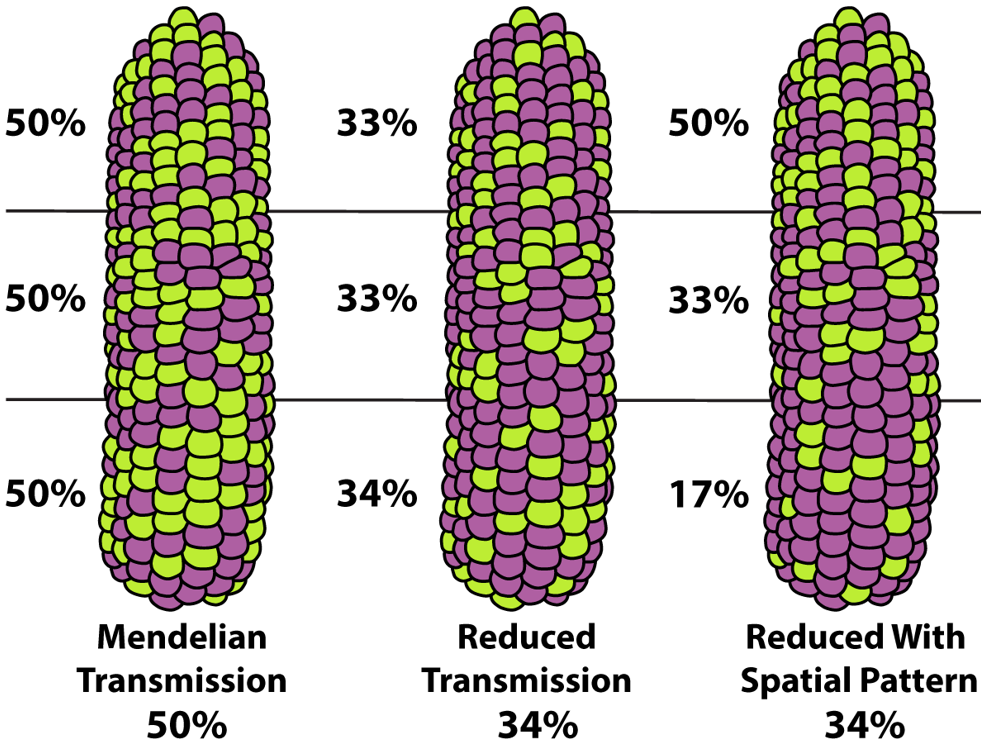
Ruggiero, D., Bang, M., Leary, M., Flieg, H., Garcia-Lamas, L., Vejlupkova, Z., Megraw, M., Jiang, D., Leiboff, S., and Fowler, J.E. "Spatial inheritance patterns across maize ears are associated with alleles that reduce pollen fitness." The Plant Journal. March 2026. [https://doi.org/10.1111/tpj.70760]
O'Hara, K., Burke, K., Ruggiero, D., and Anderson, S. "Linking Language & Thinking with Code: Computing within a Writing-Intensive Introduction to the Liberal Arts." ACM Conference on Innovation and Technology in Computer Science Education (ITiCSE). July 2017. [doi.org/10.1145/3059009.3059018]
Invited Talks
(Seminar) "Spatial inheritance patterns across maize ears are associated with alleles that reduce pollen fitness." Zeavolution Seminar. Virtual. October 2025(Short Talk) "Quantitative genetics of leaf vascular density in maize." 67th Annual Maize Genetics Meeting. St. Louis, MO. March 2025
(Lightning Talk) "Understanding leaf vascular density through quantitative genetics and high-throughput phenotyping." 66th Annual Maize Genetics Meeting, Development Premeeting. Raleigh, NC. March 2024
(Lightning Talk) "Quantitative genetics and high-throughput phenotyping of maize leaf vascular traits." 65th Annual Maize Genetics Meeting, Development Premeeting. St. Louis, MO. March 2023
(Podcast Interview) Genome Insider S3 Episode 2: Better Crops With a Pointillist Approach to Plant Genomics. DOE Joint Genome Institute. August 2022
(Short Talk) "Combinatorial barcoding for single-nucleus sequencing of developing maize leaf primordia." Plant Biology 2022, American Society of Plant Biologists. Portland, OR. July 2022
(Lightning Talk) "Single-cell genomics and high-throughput phenotyping for determining the quantitative genetics of maize leaf vascular development." 64th Annual Maize Genetics Meeting. St. Louis, MO. March 2022
Poster Presentations
FASEB Mechanisms in Plant Development Southbridge, MA. August 2025
66th Annual Maize Genetics Meeting. Raleigh, NC. March 2024
Gordon Research Conference: Single-Cell Approaches in Plant Biology. Ventura, CA. July 2023
ASPB Western Section Meeting. Tacoma, WA. April 2023
65th Annual Maize Genetics Meeting. St. Louis, MO. March 2023
2022 Plant Cell Atlas Symposium. Virtual. December 2022
ARCS Oregon Annual Scholar Event. Portland, OR. October 2022
ASPB Plant Biology 2022. Portland, OR. July 2022
64th Annual Maize Genetics Meeting. St. Louis, MO. March 2022
63rd Annual Maize Genetics Meeting. Virtual. March 2021
Fellowships
|
2024 - 2027 |
Awards
|
2024 - 2025 |
| Program providing travel and financial support, networking opportunities, and leadership development for select graduate students in the U.S. involved in research relevant to corn production | |
|
2023 |
| Travel grant to attend 2023 Gordon Research Conference for Single-Cell Approaches in Plant Biology in Ventura, CA | |
|
2023 |
| Team of three placed 2nd in multi-university hackathon challenge involving object detection for an agricultural image dataset, cash prize | |
|
2020 - 2023 |
| Support for early-career researchers of U.S. citizenship pursuing doctoral degrees in science, technology, engineering, medicine, or mathematics | |
|
2020 |
| Awarded to select newly admitted graduate students | |
|
2015 |
| Given to a premedicine or science major who maintains an interest in literature or music |
Teaching
Oregon State University, Graduate TA:BOT 101: Botany: A Human Concern - Introductory Botany for Non-Majors (In-Person)
BI 204: Introductory Biology I (Virtual)
BI 205: Introductory Biology II (Virtual)
Bard College, Undergraduate TA:
CS 143: Object-Oriented Programming with Robots (In-Person)
CS 117: Interactive Systems (In-Person)
Outreach and Leadership
|
OSU Dept. of Botany & Plant Pathology | 2022 - 2024 |
|
OSU Dept. of Botany & Plant Pathology | 2024 |
|
OSU Dept. of Botany & Plant Pathology | 2024 |
|
OSU Dept. of Botany & Plant Pathology | 2023 - 2024 |
|
OSU Dept. of Botany & Plant Pathology | 2021 - 2022 |
|
Bard College | 2015 - 2016 |